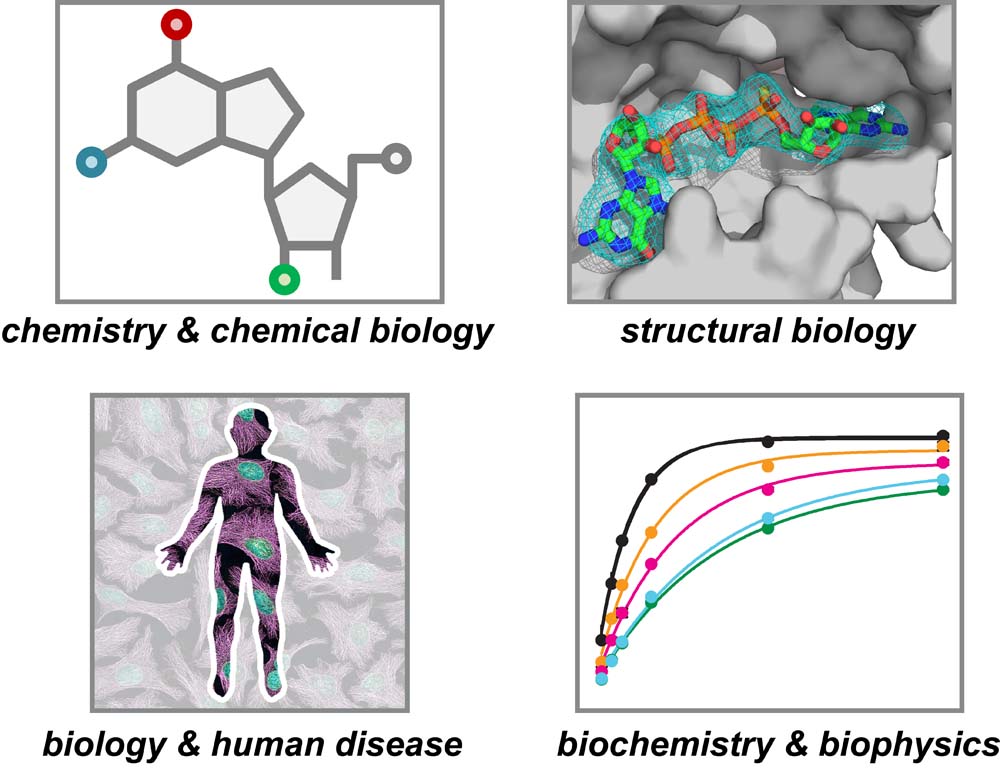
Our lab takes an interdisciplinary approach -- combining techniques and expertise from biochemistry, structural biology, biophysics, and chemistry & chemical biology -- to answer challenging questions about the regulation of RNA function and its links to human disease.
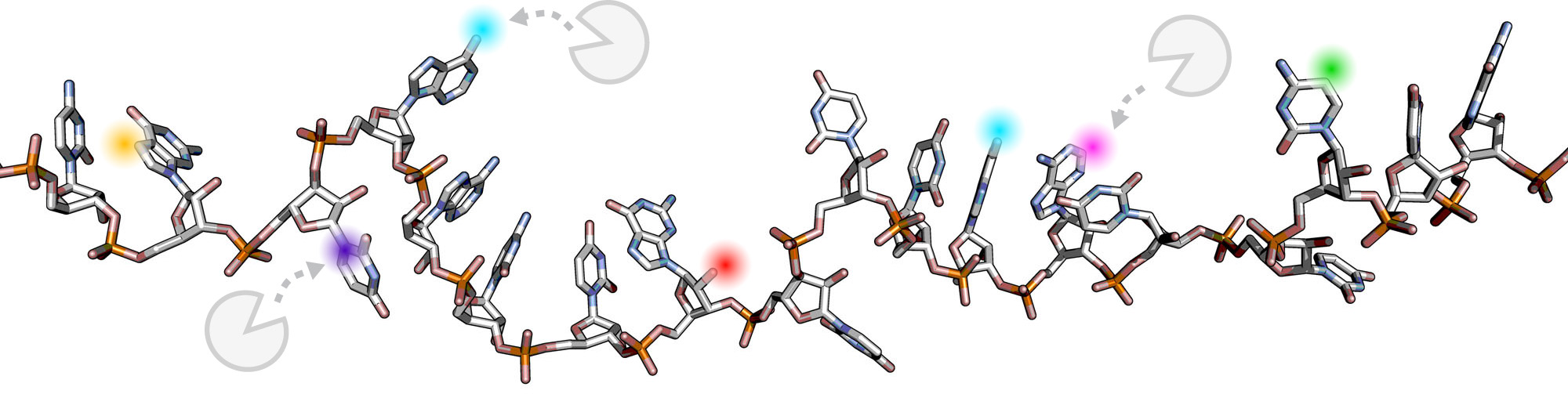
Organisms across all domains of life decorate their RNA molecules with an incredible diversity of chemical modifications. Modifications on mRNA and tRNA are critical for their function, affecting RNA structure, stability, and translational properties. Many of the proteins and enzymes that read, write, and erase these modifications are closely tied to human diseases ranging from neurological disorders to cancer to type 2 diabetes. While these proteins and pathways could be targets to treat these diseases, we lack a high-resolution, mechanistic understanding of how the cell installs, recognizes, and leverages chemical modifications on RNA. Our lab is working to understand how protein-protein and protein-nucleic acid interactions regulate chemical modifications on RNA to control gene expression and impact human disease.
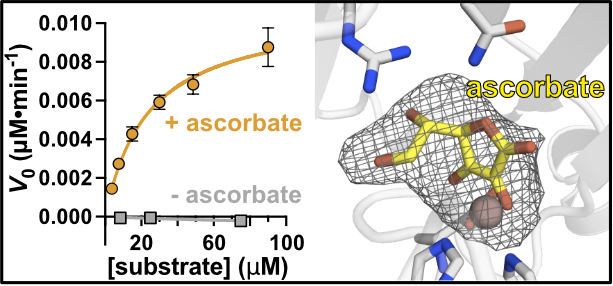
The cellular transcriptome is constantly in flux. New RNA molecules are rapidly synthesized, modified, translated, and degraded to control protein expression and respond to changes in cell environment and stress. Directly monitoring these changes in RNA composition, stability, and localization remains extremely challenging. Our lab is developing new tools to detect, map, and visualize the dynamic life of RNA inside the cell.
We’re looking for curious, enthusiastic scientists with interests in biochemistry, structural biology, chemical biology, and biophysics. Students in our lab learn and use a broad range of techniques including protein expression and purification, enzymology, RNA biochemistry, cell culture, macromolecular X-ray crystallography, mass spectrometry, NMR spectroscopy, and other biophysical assays.
The Mugridge lab is actively recruiting at all levels! We have openings for graduate students in Chemistry & Biochemistry or other programs with interests at the interface of chemistry and biology, undergrads at UD interested in doing research during the academic year and over the Summer, and postdoctoral researchers with strong backgrounds in biochemistry or structural biology. Contact Jeff directly for more information.
The Mugridge lab is committed to creating a diverse, inclusive lab space where students and researchers of all backgrounds are welcomed and supported to grow and succeed as scientists. We will actively foster a dynamic, collaborative, and open lab environment where members feel free to ask challenging questions, do innovative science, and have fun doing it!
For the latest group news, follow us on Bluesky!
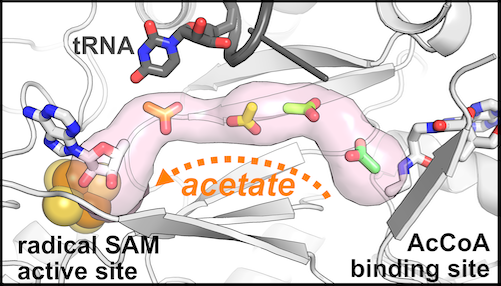
💥Latest preprint from our lab is now live! We propose a crazy(!?) new mechanism for how the radical SAM enzyme Elp3 (a subunit of the Elongator complex) modifies tRNA! #RNAsky #enzymes 🧵...
— Jeff Mugridge (@jeffmugridge.bsky.social) May 8, 2025 at 10:46 AM
[image or embed]
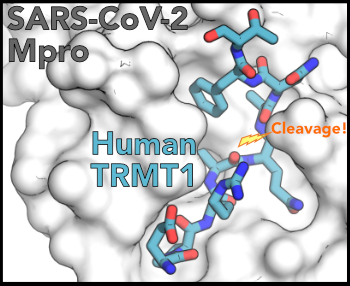
Congrats to Angel on the final version of her TRMT1 / SARS-CoV-2 Mpro story, now published in eLife! Her work reveals the structural basis for how #SARS-CoV-2 impacts #tRNA modifications! #RNAsky #RNAbiology
— Mugridge Lab (@mugridgelab.bsky.social) January 16, 2025 at 8:08 AM
[image or embed]
Big milestones - this year we said goodbye to the first two PhD grads from our lab! Now Drs. Angel D'Oliviera and Evan Geissler!👩🎓👨🎓 🎉 We also started a new lab tradition - PhD swords! ⚔️ Pictured is Angel's engraved sword with her Mpro-TRMT1 structure!
— Mugridge Lab (@mugridgelab.bsky.social) Jan 6, 2025 at 11:32 AM
[image or embed]
We finally got out and celebrated our lab's recent NSF CAREER award today with a lab bowling and laser tag trip! Pew pew! 🔫🎳
— Mugridge Lab (@mugridgelab.bsky.social) Jan 4, 2025 at 4:40 PM
[image or embed]